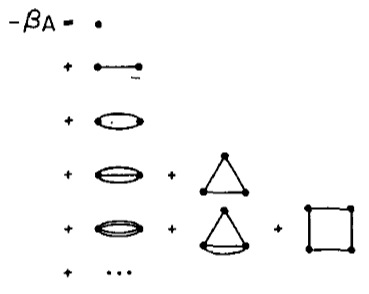
Much of condensed matter and statistical physics is concerned with the explanation of phase transitions between different forms of matter. A familiar example is water, which has a transition from the solid phase of ice to the liquid phase, and then from the liquid phase to the gaseous phase of steam, as the temperature is increased.
In fact, many different materials have all sorts of exotic phases, as you vary the temperature, pressure, or composition of the material. Constructing “phase diagrams” which show what the different phases of a material should be as a function of the varying parameters is one of the main preoccupations of physicists.
How can these phase diagrams be constructed? Condensed matter physicists tend to follow the following algorithm. First, from arguments about the microscopic physics, construct a simple model of the local interactions of the molecules making up the material. Secondly, choose some method to approximately compute the “free energy” for that simple model. Finally, find the minima of the approximate free energy as a function of the adjustable parameters like the temperature. The phase diagram can be constructed by determining which phase has the lower free energy at each point in parameter space.
If the results disagree with experiment, your model is too simple or your approximation for the free energy is not good enough, so you need to improve one or the other or both; otherwise write up your paper and submit it for publication.
To illustrate, let’s consider magnetism. Although it is less familiar than the phase transition undergone by water, magnets also have a phase transition from a magnetized “frozen” phase at low temperature to an unmagnetized “paramagnetic” phase at high temperature.
The simplest model of magnetism is the Ising model. I’ve discussed this model before; to remind you, I’ll repeat the definition of the ferromagnetic Ising model: “In this model, there are spins at each node of a lattice, that can point ‘up’ or ‘down.’ Spins like to have their neighbors point in the same direction. To compute the energy of a configuration of spins, we look at all pairs of neighboring spins, and add an energy of -1 if the two spins point in the same direction, and an energy of +1 if the two spins point in opposite directions. Boltzmann’s law tells us that each configuration should have a probability proportional to the exp(-Energy[configuration] / T), where T is the temperature.”
Of course, as defined the Ising model is a mathematical object that can and has been studied mathematically independent of any relationship to physical magnets. Alternatively, the Ising model can be simulated on a computer.
Simulations (see this applet to experiment for yourself) show that at low temperatures (and if the dimensionality of the lattice is at least 2), the lattice of spins will over time tend to align so that they point together up more than down, or they all point down more than up. The natural symmetry between up and down is “broken.” At low temperatures one will find “domains” of spins pointing in the “wrong” direction, but these domains only last temporarily.
At high temperatures, on the other hand, each spin will typically fluctuate between pointing up and down, although again domains of like-pointing spins will form. The typical time for a spin to switch from pointing up to pointing down will increase as the temperature decreases, until it diverges towards infinity (as the size of the lattice approaches infinity) as one approaches the critical temperature from above.
Intuitively, the reason for this behavior is that at low temperature, the configurations where all the spins point in the same direction have a much lower energy, and thus a much higher probability, than other configurations. At high temperatures, all the configurations start having similar probabilities, and there are many more configurations that have equal numbers of up and down spins compared to the number of aligned configurations, so one typically sees the more numerous configurations.
This balance between energetic considerations (which make the spins align) and entropic considerations (which make the spins favor the more numerous unaligned configurations) is captured by the “free energy” F, which is given by the equation F=U-TS, where U is the average energy, S is the entropy, and T is the temperature. At low temperatures, the energy dominates the free energy, while at high temperatures, the entropy dominates.
All this intuition may be helpful, but like I said, the Ising model is a mathematical object, and we should be able to find approximation methods which let us precisely calculate the critical temperature. It would also be nice to be able to precisely calculate other interesting quantities, like the magnetization as a function of the temperature, or the susceptibility (which is the derivative of the magnetization with respect to an applied field) as a function of the temperature, or the specific heat (which is the derivative of the average energy with respect to the temperature) as a function of the temperature. All these quantities can be measured for real magnets, so if we compute them mathematically, we can judge how well the Ising model explains the magnets.
This post is getting a bit long, so I’ll wait until my next post to discuss in more detail some useful methods I have worked on for systematically and precisely computing free energies, and the other related quantities which can be derived from free energies.